Creating gene libraries
by Suni Mathew
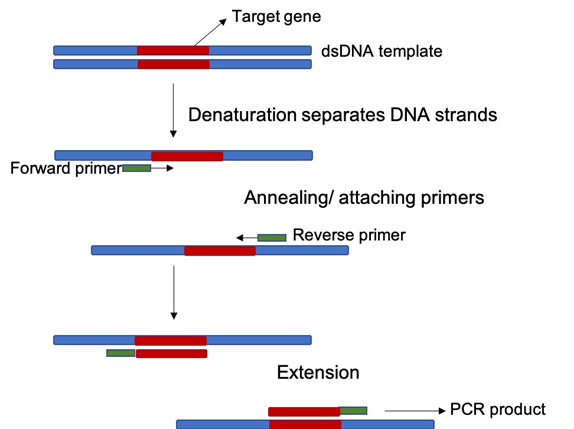
A PCR reaction is provided with a DNA template from which a gene is isolated, forward and reverse primers (or nucleotide sequences similar to the DNA sequence flanking the target gene), deoxynucleotides for synthesis of new molecules, and polymerase enzyme for gene replication. The reaction goes through three steps in a cycle – denaturation, annealing and extension – repeated several times. Hence, once the PCR reaction is done, the targeted gene is replicated exponentially, creating millions of copies compared to the starting template (Fig.1). In microbiome profiling, the 16SrRNA gene or ITS gene is isolated by PCR.
The 16SrRNA gene has variable regions V1-V9. But often two to three variable regions are targeted by PCR as the next generation sequencing machines takes an input 300-400 bp of DNA and this also reduces the complexity for data analysis afterwards. The region from V6-V8 is a favourable target region as it has been shown to identify most bacterial genera.
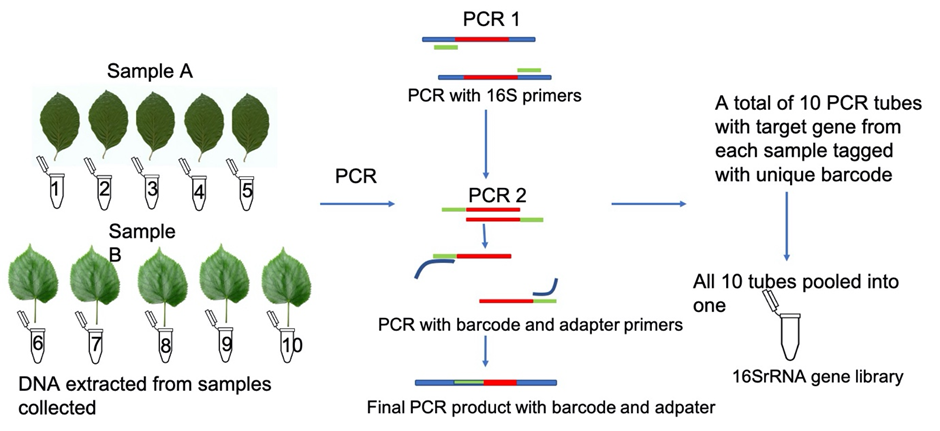
The PCR is done in a two-step approach. In the first round of PCR, the targeted gene is isolated. In the second round of PCR, the genes are targeted with special tag sequences that act as labels which will later help to track genes to their respective samples.
This can be explained with an example (Fig.2). Suppose our research question is to identify bacteria present in 2 plant samples. Each sample has 5 replicates, making the total number of samples 10. After DNA extraction, the DNA samples are subjected to first round of PCR. The first PCR products are diluted and subjected to second round of PCR. Here, the forward primers are barcodes that get tagged to the final PCR product and is thus labelled. The reverse primer is an adapter to attach the PCR products to beads in the sequencer machine. These products are pooled together, which is called a library. It is then purified and ready for sequencing.